In the previous post 5 Basic Guidelines to Carry Out a Proper EEG Recording Experiment Alejandro Riera explained how to perform a good EEG recording, with useful tips and common mistakes to avoid. Let’s say you followed the advices, and you already have a good EEG recorded. What now? Well, if you have experience with EEG, you’ll probably want to visualise the data you recorded. Or maybe even go one step further and analyse it. Perfect, that sounds like a good plan… but how? What tools should we use?
EEG software
Today we’ll try to go over the different options you have, both for visualizing and for analyzing your EEG data.
Disclaimer note: This is only a brief overview covering the software we commonly use. This post intends to be collaborative, so do not hesitate to contribute to it through your comments!
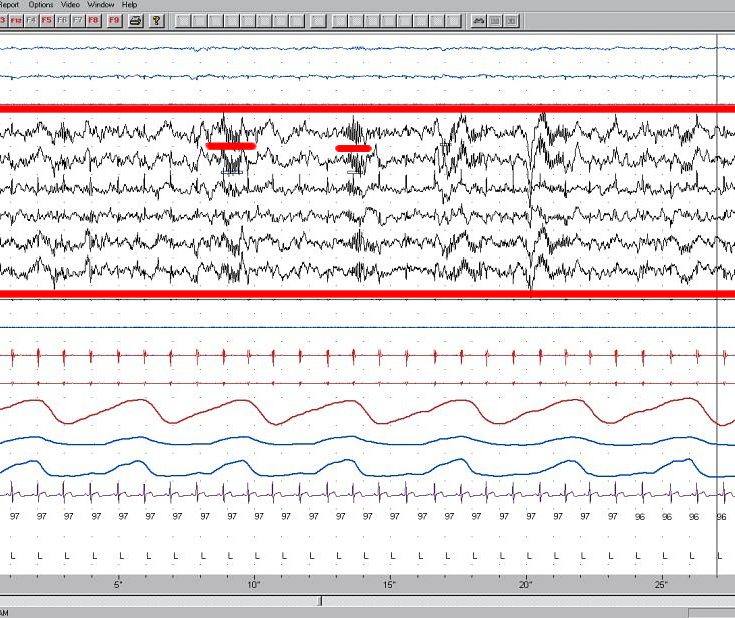
The first thing you may want to do once you finished recording your EEG data, or maybe even in between different recordings, is to visualize your data. This will enable you to visually inspect possible problems with your setup, identify artifacts like eye-blinks, or even identify artifacts from external devices. If you’re experienced, you may even be able to spot some event-related potentials on your signal, or make an estimation of the power of the signal in low frequency bands (like alpha).
Besides the proprietary applications that many commercial EEG kits bundle (like Neuroelectrics Instrument Controller, NIC, bundled with Enobio and Starstim), there are some other options of software available that you can use to visualize EEG data.
- The EDF browser supports a wide range of common EEG file-formats as input, and converts them to EDF. It offers a graphic visualization of the signal, as well as an integrated list of trigger marks present in the file. It also provides filtering functionalities, power on the frequency bands computation, as well as the possibility of down-sampling your signal. An interesting capability is that it offers the possibility to load a streaming file, so if the software bundled with your EEG package doesn’t provide any monitoring software (on-line visualization), you can use this one to monitor the signal while recording it.
- EEGLab “is an interactive Matlab toolbox for processing continuous and event-related EEG (…) data”. Besides being a processing data toolbox, it also provides visualization functionalities, both in signal plotting form for the time domain, or in a graphical-scalp drawing for the activation domain.
- Biosig is another large toolbox for Matlab (and Octave) that offers a vizualisation tool.
- Bioelectromagnetism is a “toolbox [that] has been developed to facilitate quick and easy import, visualization and measurement for ERP data.” It is specially indicated for event-related data, and is able to import/export file-formats that other software can’t (.avg, .eeg, .cnt, .tri, .3dd).
- There are other more specific software like Tempo, “an open source software for 3D visualization” providing graphical-scalp drawing for the activation domain, or EEG-Visualisation, implementing different methodologies for EEG visualization (see the webpage and the linked scientific publication there for more details).
But visualization is only the first step to take with your data. In order to work with your data and get your results, you’ll need to analyze it to detect event-related potentials, perform a time-frequency analysis, or extract some features from the signal to be used later. There is a wide range of available software, but we want to cover here the most transversal ones that we know, in the sense that it’s software that can be used in a wide range of applications.
- Matlab, of course, is a powerful tool, especially with the signal processing toolbox. Many functionalities that you may need (computation of power bands on frequency domains, detection of event-related potentials, filtering of data, feature extraction, classification, etc.) can be developed using the framework of the processing toolbox. As usual, Octave is also suitable for a lot of the work that can be developed under Matlab, with the advantage of being open source.
- If Matlab provides you the basic tools to develop, EEGLab is an open source package that already implements a vast range of different functionalities under Matlab that are commonly (and not so much) used while analyzing and processing EEG data: semi-automatic artifact removal, filtering, independent component analysis (ICA), time/frequency transforms, feature extraction, etc.
- Biosig, as we mentioned earlier, is another Matlab (and Octave) complete toolbox that provides filtering, feature extraction and classification functionalities.
- EEG-analysis-toolbox is a “MATLAB-based toolbox for exploratory analysis of EEG data. It supports both univariate analysis and multivariate pattern analysis, and can process large amounts of data in parallel.”
But if you don’t want or can’t use Matlab or Octave and you need to use a programming language, there are also software packages available that you can use to analyse EEG data in python.
- PyEEG is a Python module to extract EEG features that was initially developed for epilepsy detection, and is being upgraded. It is especially interesting for the re-projection and decomposing functionalities that it offers.
Last but not least, we would like to ask you: What software do you use for EEG analysis and which would you suggest to others? Again, this post intends to be collaborative, so don’t hesitate to contribute to it through your comments!






Hello,
According to the above it is possible to analyze from EEG: signal plotting form for the time domain, characterise in the domain of time and space in terms of voltage and frequency change, computation of power bands on frequency domains, detection of event-related potentials, filtering of data, feature extraction, classification, etc.
Is there possibility to measure speed of changeing potencials, rate of change of potentials or potential wave speed (speed per second, or per minute)?
Regards
Richard
Hi Richard,
If you’re interested in understanding more about our products, you can message info@neuroelectrics.com and we’ll get back to you the information you want.
Thank you.
Hi,
Firstly, thank you for all the valuable information. Actually, I’m a doctoral student in psychology and I need to use EEG in my research with autistic people but I don’t know the suitable software for visiualizing and analyzing the EEG data concerning this neurodevelopemental disorder. Could you propose to me some valid softwares in this situation ?
Cordially
Hi Safae,
I hope you’re still interested in these solutions, if you are, please contact info@neuroelectrics.com and we’ll share with you the information you want.
Thank you.